Products
STRUCTFAST™
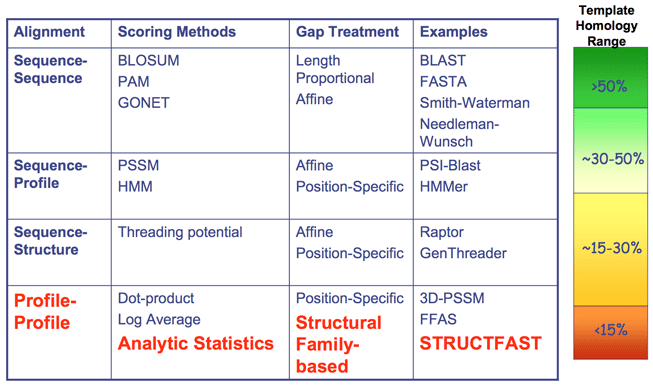
STRUCTFAST (Proteins. 2006 64:960-967) is Eidogen-Sertanty's proprietary and patent-pending protein structure determination algorithm. STRUCTFAST is the finest comparative modeling algorithm available, as demonstrated in the December 2004 CASP6 protein structure determination experiment. For comparative modeling targets, STRUCTFAST outperformed the other 47 automated servers and all but 5 of the 124 hand-modeling teams. The table below summarizes STRUCTFAST's rank by total GDT score in each of the 3 homology categories at CASP6. All GDT scores are taken from the official CASP6 website. To see a summary of the complete results, please click the header for each category.
| CASP6 Category | Rank Among All 48 Servers | Rank Among All 124 Hand Modelers |
|---|---|---|
| Comparative Modeling Easy1 | 1 | 9 |
| Comparative Modeling Hard2 | 1 | 10 |
| Comparative Modeling Combined | 1 | 6 |
The top 20 homology modeling servers at CASP6 by total GDT score included 16 hand modeling groups and 4 automated servers, including STRUCTFAST. The following table shows the number of models each of these groups placed in the top 20 according to the number of correctly aligned residues in the model (the AL0 score). Only 3 hand modeling groups in the entire CASP6 competition were able to produce high quality alignments on a more consistent basis than STRUCTFAST. Furthermore, STRUCTFAST consistently produced significantly higher quality alignments than any other automated approach.
| Group Name (Servers in Red) |
# of Models in the Top 20 |
Link to Results at the Official CASP6 Site |
|---|---|---|
| KOLINKSI-BUJNICKI | 79 | See Results |
| Jones-UCL | 69 | See Results |
| GeneSilico-Group | 60 | See Results |
| STRUCTFAST | 54 | See Results |
| BAKER | 53 | See Results |
| Ginalski | 51 | See Results |
| TOME | 51 | See Results |
| Skolnick-Zhang | 50 | See Results |
| CBRC-3D | 38 | See Results |
| FISCHER | 37 | See Results |
| CHIMERA | 34 | See Results |
| SAM-T04-hand | 29 | See Results |
| SBC | 28 | See Results |
| Sternberg | 27 | See Results |
| CAFASP-Consensus | 26 | See Results |
| zhousp3 | 23 | See Results |
| ZHOUSPARKS2 | 23 | See Results |
| ACE | 23 | See Results |
| SBC-Pmodeller5 | 19 | See Results |
CASP6 Category Definitions:
- Comparative Modeling Easy: Low to moderate homology prediction targets where BLAST finds a homolog in the PDB.
- Comparative Modeling Hard: Low homology prediction targets where BLAST cannot find a homolog in the PDB , but PSI-BLAST can.
