Products
Eidogen Visualization Environment™ (EVE™)
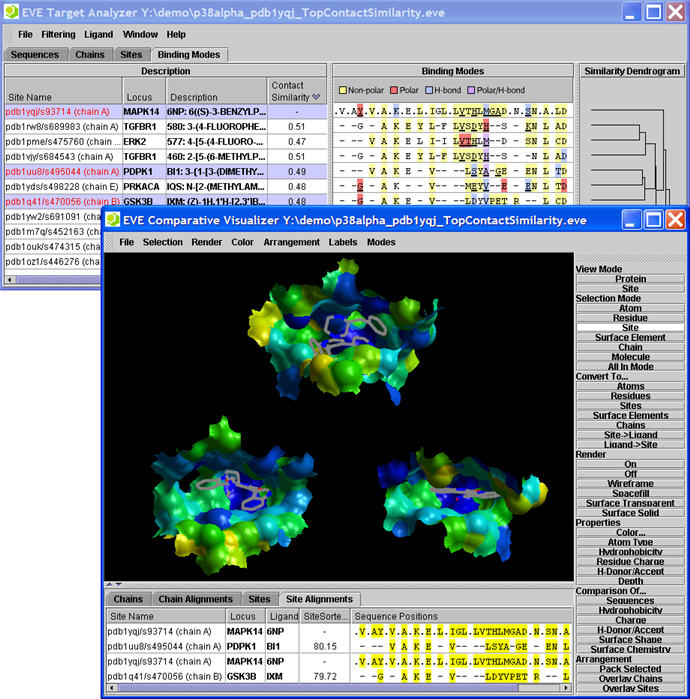
The EVE™ analysis tool allows users to visualize and compare the small molecule binding sites of targets of interest in a matter of seconds. With EVE's powerful yet easy-to-use Java3D©-driven interface, both expert and non-expert users alike can quickly analyze and compare important binding site properties and ligand interactions, dramatically enhancing the efficiency with which structure-based selectivity, binding mode analysis, and lead optimization studies can be carried out.
Key features: Eidogen Visualization Environment
- Easy visualization of Sequence, Structure, Site, and Site-Ligand Interaction similarity relationships.
- Site-Ligand Contact (SLiC) analysis for comparison of ligand binding modes
- Compare ligand-bound structures based on site-ligand interaction fingerprints.
- Understand ligand selectivity across multiple targets.
- Create composite Site-Ligand Contact fingerprints (cSLiCs) from a set of co-crystal or docked compounds
- Import docking/virtual screening results to rescore docked poses based on SLiC/cSLiC scores
- Customizable parameters for tuning SLiC/cSLiC scoring for ligand interactions important for compound or target classes of interest
- LigandCross for creation of novel ligand scaffolds from structural data.
- Fast, scalable method leverages existing co-crystal or docked structures to generate new ligands from fragments known to bind to the target of interest.
- Create all possible combinations of known binding fragments from a set of co-crystallized or docked compounds and export 3D structures
- Example case - Kinases with DFG-out co-complexes
- Easily clone ligands into multiple binding sites to understand basis for selectivity and cross-reactivity
- Perform side-by-side comparisons and overlays of whole protein structures or individual binding sites using pre-computed alignments.
- Easy mapping and annotation of important binding site properties such as hydrophobicity, charge, hydrogen-bonding capability and pocket depth.
- Locus annotation for quick identification of targets of interest.
- Visualization of entire Project Files rather than individual structures enables easy sharing of project data between scientists.
- Complete macro language and batch-mode compatibility for carrying out customized, large-scale analyses.